-Search query
-Search result
Showing all 38 items for (author: sansom & msp)

EMDB-15641: 
Structure of the Native Chemotaxis Core Signalling Complex from E-gene lysed E. coli cells.
Method: subtomogram averaging / : Qin Z, Zhang P

EMDB-15642: 
Native Chemotaxis Core Signalling Complex from E. coli, with focused alignment on the baseplate (CheA-CheW)
Method: subtomogram averaging / : Qin Z, Zhang P
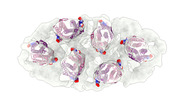
EMDB-15643: 
Native Chemotaxis Core Signalling Complex from E. coli, Focused alignment on ligand binding domain
Method: subtomogram averaging / : Qin Z, Zhang P

EMDB-23700: 
Full length alpha1 Glycine receptor in presence of 32uM Tetrahydrocannabinol
Method: single particle / : Kumar A, Chakrapani S

EMDB-23701: 
Full length alpha1 Glycine receptor in presence of 0.1mM Glycine
Method: single particle / : Kumar A, Chakrapani S

EMDB-23702: 
Full length alpha1 Glycine receptor in presence of 0.1mM Glycine and 32uM Tetrahydrocannabinol
Method: single particle / : Kumar A, Chakrapani S

EMDB-23703: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine
Method: single particle / : Kumar A, Chakrapani S

EMDB-23704: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine and 32uM Tetrahydrocannabinol State 1
Method: single particle / : Kumar A, Chakrapani S

EMDB-23705: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine and 32uM Tetrahydrocannabinol State 2
Method: single particle / : Kumar A, Chakrapani S

EMDB-23706: 
Full length alpha1 Glycine receptor in presence of 1mM Glycine and 32uM Tetrahydrocannabinol State 3
Method: single particle / : Kumar A, Chakrapani S

EMDB-13764: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Composite Map)
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-14578: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Consensus Map)
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-24186: 
Composite Structure of Mechanosensitive Ion Channel Flycatcher1 in GDN
Method: single particle / : Jojoa-Cruz S, Saotome K, Lee WH, Patapoutian A, Ward AB

EMDB-24187: 
Structure of Mechanosensitive Ion Channel Flycatcher1 in GDN
Method: single particle / : Jojoa-Cruz S, Saotome K, Lee WH, Patapoutian A, Ward AB

EMDB-24188: 
Structure of Mechanosensitive Ion Channel Flycatcher1 Protomer in 'Down' conformation in GDN
Method: single particle / : Jojoa-Cruz S, Saotome K, Lee WH, Patapoutian A, Ward AB

EMDB-24189: 
Structure of Mechanosensitive Ion Channel Flycatcher1 Protomer in 'Up' conformation in GDN
Method: single particle / : Jojoa-Cruz S, Saotome K, Lee WH, Patapoutian A, Ward AB

EMDB-13860: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to IMP-1575
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-13841: 
Focused refinement of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-13842: 
Focused refinement of Hedgehog acyltransferase (HHAT) in complex with megabody 177 - megabody core
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-11557: 
YnaI
Method: single particle / : Flegler VJ, Rasmussen A

EMDB-11560: 
YnaI in an open-like conformation
Method: single particle / : Flegler VJ, Rasmussen A

EMDB-11556: 
small conductance mechanosensitive channel YnaI mutant G149A/G152A
Method: single particle / : Flegler VJ, Rasmussen A, Rao S, Wu N, Zenobi R, Sansom MSP, Hedrich R, Rasmussen T, Boettcher B

EMDB-11558: 
mechanosensitive channel YnaI in amphipol A8-35
Method: single particle / : Flegler VJ, Rasmussen A, Rao S, Wu N, Zenobi R, Sansom MSP, Hedrich R, Rasmussen T, Boettcher B

EMDB-11559: 
mechanosensitive channel YnaI in a closed-like conformation
Method: single particle / : Flegler VJ, Rasmussen A, Rao S, Wu N, Zenobi R, Sansom MSP, Hedrich R, Rasmussen T, Boettcher B

EMDB-11561: 
mechanosensitive channel YnaI in an intermediate conformation
Method: single particle / : Flegler VJ, Rasmussen A, Rao S, Wu N, Zenobi R, Sansom MSP, Hedrich R, Rasmussen T, Boettcher B

EMDB-11629: 
small conductance mechanosensitive channel YbiO
Method: single particle / : Flegler VJ, Rasmussen A

EMDB-20714: 
Full length Glycine receptor reconstituted in lipid nanodisc in Apo/Resting conformation
Method: single particle / : Kumar A, Basak S, Chakrapani S

EMDB-20715: 
Full length Glycine receptor reconstituted in lipid nanodisc in Gly-bound desensitized conformation
Method: single particle / : Kumar A, Basak S, Chakrapani S

EMDB-20731: 
Full length Glycine receptor reconstituted in lipid nanodisc in Gly/PTX-bound open/blocked conformation
Method: single particle / : Kumar A, Basak S, Chakrapani S

EMDB-21234: 
Full length Glycine receptor reconstituted in lipid nanodisc in Gly/IVM-conformation (State-1)
Method: single particle / : Kumar A, Basak S, Chakrapani S

EMDB-21236: 
Full length Glycine receptor reconstituted in lipid nanodisc in Gly/IVM-conformation (State-2)
Method: single particle / : Kumar A, Basak S, Chakrapani S

EMDB-21237: 
Full length Glycine receptor reconstituted in lipid nanodisc in Gly/IVM-conformation (State-3)
Method: single particle / : Kumar A, Basak S, Chakrapani S

EMDB-10418: 
CryoEM structure of human polycystin-2/PKD2 in UDM supplemented with PI(4,5)P2
Method: single particle / : Wang Q, Pike ACW, Grieben M, Baronina A, Nasrallah C, Shintre C, Edwards AM, Arrowsmith CH, Bountra C, Carpenter EP, Structural Genomics Consortium (SGC)

EMDB-10419: 
CryoEM structure of human polycystin-2/PKD2 in UDM supplemented with PI(3,5)P2
Method: single particle / : Wang Q, Pike ACW, Grieben M, Baronina A, Nasrallah C, Shintre C, Edwards AM, Arrowsmith CH, Bountra C, Carpenter EP, Structural Genomics Consortium (SGC)

EMDB-7882: 
Full-length 5-HT3A receptor in a serotonin-bound conformation- State 1
Method: single particle / : Basak S, Chakrapani S

EMDB-7883: 
Full-length 5-HT3A receptor in a serotonin-bound conformation- State 2
Method: single particle / : Basak S, Chakrapani S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
